- Blog
- Summon quality vs skeleton level
- Sonic racing volume reset
- Super photocut pro
- Jaccard similarity
- Download screen recorder
- Perfect diet tracker for ios wont install
- Razorlame mp3 encoder
- C- mirabai olympic games tokyo 2020
- Black podengo
- Look at specific amino acids jmol
- Tiltshift canon
- War thunder japanese pacific campaign
- Linkedin daily active users
- Online athens police blotter

Rotate the image and zoom to examine the key contacts. You should see new atoms from the protein appear-the ligand is covalently bound to some, and others are in vaan der Waals contact. Then, turn on Wireframe and Spacefill for the selected atoms in the Appearance panelĪnd color them CPK using the Color panel.
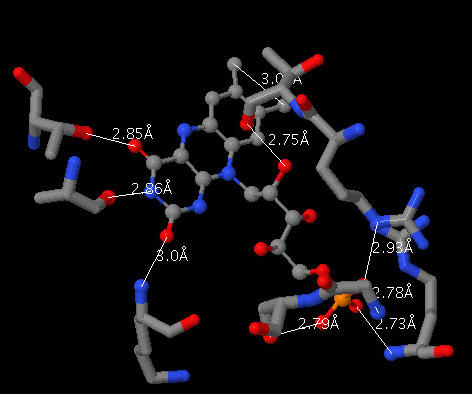
In the Custom Selector field of the Advanced Select panel, type Within 6.0 Å of atom number 2087, the ligand's phosphorus atom.
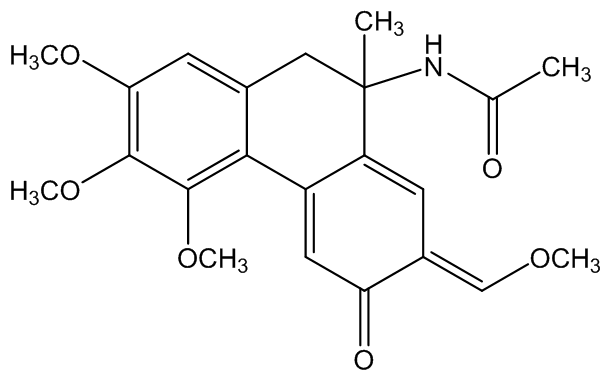
The custom selector "within" can help you explore how a ligand isīound to a protein by selecting only atoms that are in close proximity to the ligand. Now let's determine the atom(s) in the protein that make contact The geometry at the central phosphorus atom is best described as. Turn on Wireframe and Spacefill, then use the Color Panel to color the bonds & atoms " CPK."
#Look at specific amino acids jmol code#
Using the Advanced Select panel (Tools → Advanced Select), select residues with the three-letter code "CYS." Use the Appearance panel to Residues as balls and sticks and color their atoms by element To do this we will show the cysteine (CYS) Click "None" to clear your selection to see the colors clearly. Secondary structural elements can now be distinguished by their colors. Pull up the Color panel (Look & Feel → Color), select "Cartoon," and color the cartoon "Structure." Notice that the "Cartoon" to show the chain as a cartoon. Then, pull up the Appearance panel (Look & Feel → Appearance) and turn off Wireframe and Spacefill. The select mode bar below the Jmol window and clicking any atom in the polypeptide. Select the entire protein by clicking "Chain" in Let's begin by cartooning the protein and coloring it according to the secondary structural elements present. You should see an image like the one at right appear.Įxploring 4TGL. You can obtain 4TGL by navigating to the Jmol Interface and clicking "File → PDB Entry." Type "4TGL" into the text field that appears, and press Enter. This assignment assumes you are familiar with the Jmol Interface at the level of the first Jmol quiz. As you build the model, you will see the primary structure, secondary structure and a simplified version of the tertiary structure of tRNA.Last Command: Getting Started.
#Look at specific amino acids jmol pdf#
Use this PDF to build a paper model of the tRNA. To learn more, see the Molecule of the Month features on tRNA and Ribosome. Each tRNA molecule binds to a specific amino acid on the acceptor arm, recognizes its corresponding codon in the mRNA through the anticodon loop region, and delivers the amino acid to a growing peptide chain in the ribosome for protein synthesis. Explore the atomic structure of tRNA - Interactive display of the atomic model of tRNA and details about its structureĭuring protein synthesis in cells, transfer RNA (tRNA) "translates" the genetic code in the messenger RNA (mRNA) into the language of proteins.Build a paper model of tRNA - Template and instructions for making the paper model.What is tRNA? - An introduction to the molecule.You can use an online interactive display of tRNA's atomic model and the paper model to learn about the structure and function of this molecule. In this activity you will make a paper model of a biological molecule - tRNA.
